Physiologic Monitoring and Countermeasures Applied to Humans in Space and Extreme Environments
Use of Complex Sensor Arrays, Convergent Variable Detection, and Mathematical Modeling
Location: NASA Ames Research Center, Psychophysiology Research Laboratory
Laboratory Directors: Cowings, P, Toscano, WB
Fellowship-Molecular Lead: Schmidt, MA (Previous Work)
The Psychophysiology Research Laboratory (PRL) at NASA Ames Research Center has been engaged for several decades in the simultaneous measurement of multiple physiologic variables in humans in extreme environments. This work has been coupled with the development of countermeasure solutions for these same environments. Spaceflight is the primary focus of this work, but the work also extends to military field operations and clinical medicine.
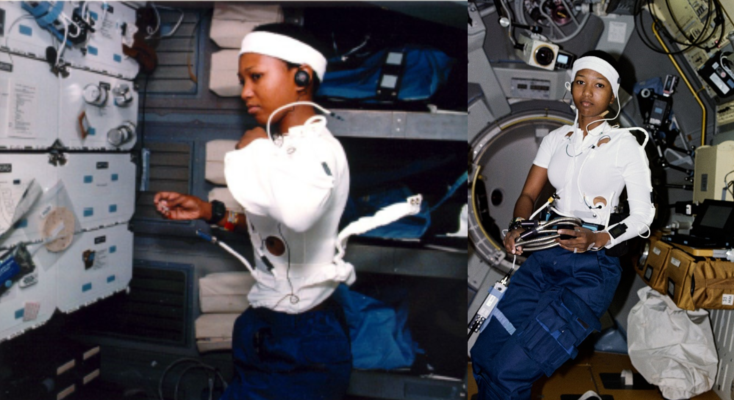
During this work, the PRL has developed two notable technologies. The first is the Ambulatory Autogenic Feedback System-2, which is a garment containing the sensors needed to monitor selected physiologic signals in a field setting (Figure 4). The second is a training technique called Autogenic-Feedback Training Exercise (AFTE), which is the countermeasure approach used to improve tolerance and performance in extreme environments. This is based on the real-time assessment of up to 20 different physiologic measures (Table 1) for assessing the response in extreme environments and for use in training the individual to self-regulate biological function and performance (Figures 1-3). Both technologies have been deployed on astronauts in space environments and in novel performance contexts, such as the space shuttle and the Naval Aerospace Medical Institute (Figure 5).
GENERAL MEASURES
- Physiologic: autonomic nervous system, central nervous system
- Performance: cognitive, perceptual, neuromotor
- Subjective: mood, symptoms
Molecular Dynamics Beneath the Physiologic Phenotype
This extreme physiology work was also part of our (Schmidt MA et al.) two decades-long effort (and ongoing) to better describe the dynamic interaction between molecular networks and the physiologic phenotype we typically measure. It is evident that there is a progression wherein molecular networks underlie the physiologic, morphologic, and behavioral phenotypes. As such, when sensor systems are being deployed in the measurement of physiologic signals, it is the molecular networks that govern these signals. Of course, there is constant modulation of the molecular dynamics and the physiologic signals. But to better understand the dynamics of human performance and human disease, we must develop greater sophistication in describing these relationships. In short, we have been examining this linkage between molecular networks, physiologic phenotype, clinical phenotype, and performance phenotype. The physiologic signals are often highly enriched, meaning they are ripe with continuous variables that change over time. Simple molecular measures (targeted) in such enriched physiologic signal environments are not wholly informative. But the evolution of integrated omics methods (untargeted genomics, transcriptomics, proteomics, metabolomics, etc.) allows for enriched molecular signals that can be analyzed against the enriched physiologic signals. Data reduction techniques and multivariate modeling tools applied to these high dimensional data sets affords us the ability to better describe these patterns and help to identify potential actionable countermeasure targets (or therapeutic targets) in the molecular networks.
Table 1 Physiologic Measures The primary physiologic measures are shown below. These are often coupled with EEG measurements when the environmental or study conditions permit.
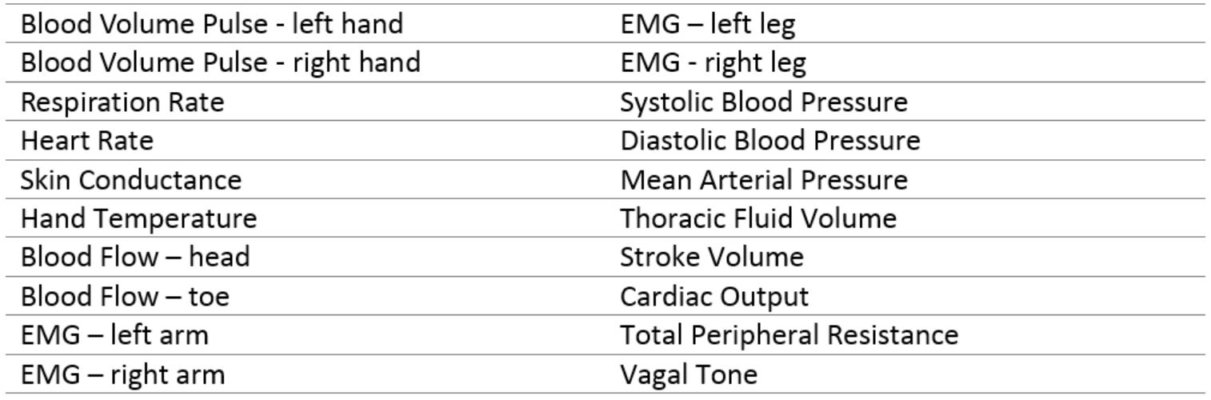
Figure 1. Digital Display of 20+ Physiologic Measures Derived from Sensor System All AFTE approaches utilize both digital and analog displays on a computer monitor. The monitors display real-time results of each of the physiologic measures being captured by the sensors. The facilitator/clinician can display only one or up to all 20 of these measures simultaneously (in both digital and analog form), while training the subject in self-regulation of these physiologic parameters. (Image Credit: Psychophysiology Research Laboratory, NASA)

Figure 2. NASA Autogenic Clinical Laboratory System: Facilitator Suite (Image Credit: Psychophysiology Research Laboratory, NASA)

Figure 3. NASA Autogenic Clinical Laboratory System: Training Suite (Image Credit: Psychophysiology Research Laboratory, NASA)
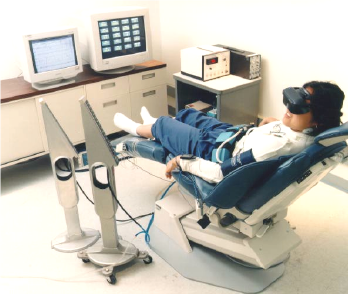
Figure 4. NASA Ambulatory Autogenic Feedback System-2 A schematic representation of signals captured by the ambulatory garment system developed by Cowings & Toscano (PRL). This system has been deployed in space to provide real-time measures of a selected number of physiologic measures. The astronaut is able to monitor function and engage in real-time countermeasure training, while in space. This is intended to reinforce the training provided on Earth. (Image Credit: Psychophysiology Research Laboratory, NASA)

Figure 5. Astronaut Mae Jemison, M.D. on the space shuttle Endeavor (STS-47), wearing the physiologic monitoring vest (Autogenic Feedback System-2) developed by the NASA Ames Psychophysiology Laboratory (Cowings & Toscano; Image Credit: NASA)

